1.1.3 Suffix Trees and Arrays
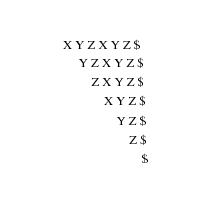
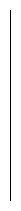
INPUT OUTPUT
Input Description:
A reference string S.
Problem:
Build a data structure for quickly finding all places where
an arbitrary query string q is a substring of S.
Excerpt from
The Algorithm Design Manual:
Suffix trees and arrays are phenomenally useful data structures for solving
string problems elegantly and efficiently.
If you need to speed up a string processing algorithm from
O(n^2) to linear time, proper use of suffix trees is quite likely the
answer.
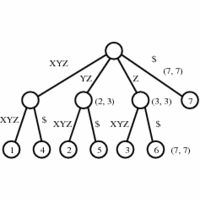
In its simplest instantiation, a suffix tree is simply a trie
of the n strings that are suffixes of an n-character string S.
A trie is a tree structure, where each node represents one character,
and the root represents the null string.
Thus each path from the root represents a string, described by the
characters labeling the nodes traversed.
Any finite set of words defines a trie, and two words with
common prefixes will branch off from each other at the first distinguishing
character.
Each leaf represents the end of a string.
Tries are useful for testing whether a given query string q
is in the set.
Starting with the first character of q, we traverse the trie along the
branch defined by the next character of q.
If this branch does not exist in the trie, then q cannot be one of the set
of strings.
Otherwise we find q in |q| character comparisons regardless
of how many strings
are in the trie.
Tries are very simple to build (repeatedly insert new strings)
and very fast to search, although they can be
expensive in terms of memory.
A suffix tree is simply a trie of all the proper suffixes
of S.
The suffix tree enables you to quickly test whether q is a substring of
S, because any substring of S is the prefix of some suffix (got it?).
The search time is again linear in the length of q.
Recommended Books
Related Links
white page links and source
Related Problems
This page last modified on 2008-07-10
.
www.algorist.com



 Data Structures and Algorithms
Data Structures and Algorithms




